Contents
- CPWC Double Adaptive Beamforming - Explicit Processing (No Pipeline)
- Download and load channel data
- Define scan
- Create apodization objects
- Create midprocess DAS object
- Configure Capon Minimum Variance beamformers
- =========================================================================
- IMAGE 1: DAS on RX -> DAS on TX : Conventional
- =========================================================================
- =========================================================================
- IMAGE 2: MV on RX -> DAS on TX
- =========================================================================
- =========================================================================
- IMAGE 3: DAS on RX -> MV on TX
- =========================================================================
- =========================================================================
- IMAGE 4: MV on RX -> MV on TX : Double Adaptive
- =========================================================================
- =========================================================================
- IMAGE 5: DAS on TX -> MV on RX
- =========================================================================
- Add coherence in the mix
- =========================================================================
- Display Results
- =========================================================================
- Plot single image
- Plot Figure - All 5 images
- Plot timing bar chart
- Save figures
- Compare beamformed data
- Plot timing bar chart
- =========================================================================
- gCNR Analysis
- =========================================================================
- =========================================================================
- FWHM / 6dB Resolution Analysis
- =========================================================================
- =========================================================================
- Lateral Profile Plot
- =========================================================================
- =========================================================================
- Combined Bar Charts Figure
- =========================================================================
- =========================================================================
- Summary Table
- =========================================================================
- =========================================================================
- Local Functions
- =========================================================================
CPWC Double Adaptive Beamforming - Explicit Processing (No Pipeline)
This script performs double adaptive beamforming using Capon Minimum Variance (MV) beamforming. It uses explicit midprocess/postprocess calls instead of the pipeline feature - the conventional way of doing processing in USTB.
Five different beamforming configurations are compared: 1. DAS on RX -> DAS on TX (Conventional) 2. MV on RX -> DAS on TX 3. DAS on RX -> MV on TX 4. MV on RX -> MV on TX (Double Adaptive) 5. DAS on TX -> MV on RX
Author: Ole Marius Hoel Rindal olemarius@olemarius.net
clear all; close all;
Download and load channel data
url = tools.zenodo_dataset_files_base(); filename = 'PICMUS_numerical_calib_v2.uff'; tools.download(filename, url, data_path); channel_data = uff.channel_data(); channel_data.read([data_path, filesep, filename], '/channel_data');
USTB download tool File: PICMUS_numerical_calib_v2.uff URL: https://zenodo.org/records/19550715/files/PICMUS_n umerical_calib_v2.uff Path: C:\Repositories\ustb\data Downloading 2 / 2 MB...done! UFF: reading /channel_data [uff.channel_data] UFF: reading sequence [uff.wave] [====================] 100%
Define scan
scan = uff.linear_scan('x_axis', linspace(-20e-3, 20e-3, 256).', ... 'z_axis', linspace(10e-3, 50e-3, 256).'); %scan = uff.linear_scan('x_axis', linspace(-20e-3, 20e-3, 768).', ... % 'z_axis', linspace(10e-3, 50e-3, 512).'); %scan=uff.linear_scan('x_axis',linspace(0e-3,15e-3,256).', 'z_axis', linspace(23e-3,27e-3,256).');
Create apodization objects
Transmit apodization
transmit_apodization = uff.apodization(); transmit_apodization.probe = channel_data.probe; transmit_apodization.sequence = channel_data.sequence; transmit_apodization.focus = scan; transmit_apodization.window = uff.window.tukey75; transmit_apodization.minimum_aperture = [3.07000e-03 3.07000e-03]; transmit_apodization.f_number = 1.75; transmit_apodization_with_window = transmit_apodization; % Receive apodization receive_apodization = uff.apodization(); receive_apodization.probe = channel_data.probe; receive_apodization.focus = scan; receive_apodization.window = uff.window.tukey25; receive_apodization.f_number = 1.75; % Receive apodization for MV on tx receive_apodization_none_window = uff.apodization(); receive_apodization_none_window.probe = channel_data.probe; receive_apodization_none_window.focus = scan; receive_apodization_none_window.window = uff.window.none; % Pre-compute apodization data (so it doesn't affect timing) transmit_apodization.data; receive_apodization.data; receive_apodization_none_window.data;
Create midprocess DAS object
das = midprocess.das(); das.channel_data = channel_data; das.scan = scan; das.transmit_apodization = transmit_apodization; das.receive_apodization = receive_apodization;
Configure Capon Minimum Variance beamformers
Receive adaptive (Capon MV)
adapt_rx = postprocess.capon_minimum_variance(); adapt_rx.dimension = dimension.receive; adapt_rx.scan = scan; adapt_rx.channel_data = channel_data; adapt_rx.K_in_lambda = 1.5; adapt_rx.L_elements = floor(channel_data.probe.N_elements/2); adapt_rx.regCoef = 1/100; adapt_rx.transmit_apodization = transmit_apodization; adapt_rx.receive_apodization = receive_apodization; % Transmit adaptive (Capon MV) adapt_tx = postprocess.capon_minimum_variance(); adapt_tx.dimension = dimension.transmit; adapt_tx.scan = scan; adapt_tx.channel_data = channel_data; adapt_tx.K_in_lambda = 1.5; adapt_tx.L_elements = channel_data.probe.N_elements; adapt_tx.regCoef = 1/100; adapt_tx.transmit_apodization = transmit_apodization; adapt_tx.receive_apodization = receive_apodization; % Display settings compression = 'log'; dynamic_range = 60;
=========================================================================
IMAGE 1: DAS on RX -> DAS on TX : Conventional
=========================================================================
fprintf('\n=== Processing Image 1: Conventional DAS ===\n'); das.transmit_apodization = transmit_apodization; transmit_apodization.data; das.dimension = dimension.both; tic(); img{1} = das.go(); time{1} = toc(); label{1} = 'DASonRX->DASonTX'; fprintf('Completed in %.2f seconds.\n', time{1}); img{1}.plot()
=== Processing Image 1: Conventional DAS ===
USTB MEX C beamformer...Completed in 0.08 seconds.
Completed in 0.09 seconds.
ans =
Figure (1) with properties:
Number: 1
Name: ''
Color: [1 1 1]
Position: [323 225 1061 639]
Units: 'pixels'
Use GET to show all properties

=========================================================================
IMAGE 2: MV on RX -> DAS on TX
=========================================================================
fprintf('\n=== Processing Image 2: MV on RX -> DAS on TX ===\n'); % Step 1: DAS with no compounding (dimension.none) das.dimension = dimension.none; das.transmit_apodization = transmit_apodization; % Update adapt_rx parameters adapt_rx.L_elements = floor(2*channel_data.probe.N_elements/3); adapt_rx.regCoef = 1/100; tic(); b_data_das = das.go(); % Step 2: Apply adaptive RX beamforming adapt_rx.input = b_data_das; b_data_adapt_rx = adapt_rx.go(); % % Step 3: Coherent compounding on TX dimension das_tx = postprocess.coherent_compounding(); das_tx.dimension = dimension.transmit; das_tx.receive_apodization = receive_apodization; das_tx.transmit_apodization = transmit_apodization_with_window; das_tx.input = b_data_adapt_rx; img{2} = das_tx.go(); time{2} = toc(); label{2} = 'MVonRX->DASonTX'; fprintf('Completed in %.2f seconds.\n', time{2}); img{2}.plot()
=== Processing Image 2: MV on RX -> DAS on TX ===
USTB MEX C beamformer...Completed in 0.10 seconds.
Completed in 95.94 seconds.
ans =
Figure (2) with properties:
Number: 2
Name: ''
Color: [1 1 1]
Position: [323 225 1061 639]
Units: 'pixels'
Use GET to show all properties


=========================================================================
IMAGE 3: DAS on RX -> MV on TX
=========================================================================
fprintf('\n=== Processing Image 3: DAS on RX -> MV on TX ===\n'); % Update adapt_tx parameters adapt_tx.L_elements = floor(channel_data.N_waves/3); adapt_tx.regCoef = 1/100; adapt_tx.receive_apodization = receive_apodization; % Step 1: DAS on receive only das.dimension = dimension.receive; das.receive_apodization = receive_apodization; tic(); b_data_das = das.go(); % Step 2: Apply adaptive TX beamforming adapt_tx.input = b_data_das; img{3} = adapt_tx.go(); time{3} = toc(); label{3} = 'DASonRX->MVonTX'; fprintf('Completed in %.2f seconds.\n', time{3});
=== Processing Image 3: DAS on RX -> MV on TX === USTB MEX C beamformer...Completed in 0.09 seconds. Completed in 7.55 seconds.
=========================================================================
IMAGE 4: MV on RX -> MV on TX : Double Adaptive
=========================================================================
fprintf('\n=== Processing Image 4: MV on RX -> MV on TX (Double Adaptive) ===\n'); % Update parameters adapt_tx.L_elements = floor(channel_data.N_waves/3); adapt_tx.regCoef = 1/100; % Step 1: DAS with no compounding das.dimension = dimension.none; tic(); b_data_das = das.go(); % Step 2: Apply adaptive RX beamforming adapt_rx.input = b_data_das; b_data_adapt_rx = adapt_rx.go(); % Step 3: Apply adaptive TX beamforming adapt_tx.input = b_data_adapt_rx; img{4} = adapt_tx.go(); time{4} = toc(); label{4} = 'MVonRX->MVonTX'; fprintf('Completed in %.2f seconds.\n', time{4});
=== Processing Image 4: MV on RX -> MV on TX (Double Adaptive) === USTB MEX C beamformer...Completed in 0.10 seconds. Completed in 102.36 seconds.
=========================================================================
IMAGE 5: DAS on TX -> MV on RX
=========================================================================
fprintf('\n=== Processing Image 5: DAS on TX -> MV on RX ===\n'); % Update adapt_rx parameters adapt_rx.L_elements = floor(channel_data.probe.N_elements/2); adapt_rx.regCoef = 1/100; % Step 1: DAS on transmit only das.dimension = dimension.transmit; das.receive_apodization = receive_apodization_none_window tic(); b_data_das = das.go(); % Step 2: Apply adaptive RX beamforming adapt_rx.input = b_data_das; img{5} = adapt_rx.go(); time{5} = toc(); label{5} = 'DASonTX->MVonRX'; fprintf('Completed in %.2f seconds.\n', time{5});
=== Processing Image 5: DAS on TX -> MV on RX ===
das =
das with properties:
dimension: transmit
code: mex
gpu_id: 0
spherical_transmit_delay_model: hybrid
pw_margin: 1.0000e-03
lens_delay: 0
blending_power: 0.5000
transmit_delay: [65536×5 single]
receive_delay: [65536×128 single]
elapsed_time: 0.0981
channel_data: [1×1 uff.channel_data]
scan: [1×1 uff.linear_scan]
receive_apodization: [1×1 uff.apodization]
transmit_apodization: [1×1 uff.apodization]
beamformed_data: [1×1 uff.beamformed_data]
name: 'USTB DAS General Beamformer'
reference: 'www.ustb.no'
implemented_by: {1×3 cell}
version: 'v1.1.0'
USTB MEX C beamformer...Completed in 0.22 seconds.
Completed in 32.48 seconds.
Add coherence in the mix
cf = postprocess.coherence_factor; cf.input = b_data_das; b_data_cf = cf.go(); b_data_cf.plot([], 'Coherence Factor', [], 'abs'); caxis([0 1]); cf.CF.plot([], 'Coherence Factor', [], 'abs'); img_cf_temp = uff.beamformed_data(img{5}); img_cf_temp.data = img{5}.data.*cf.CF.data.^0.3; img_cf_temp.plot([], 'MVonRX->DASonTX with CF', dynamic_range, compression);
Warning: Not enough waves to compute factor. Changing dimension to dimension.receive




=========================================================================
Display Results
=========================================================================
fprintf('\n=== Processing Complete ===\n'); fprintf('Total processing times:\n'); for i = 1:5 fprintf(' %s: %.2f seconds\n', label{i}, time{i}); end
=== Processing Complete === Total processing times: DASonRX->DASonTX: 0.09 seconds MVonRX->DASonTX: 95.94 seconds DASonRX->MVonTX: 7.55 seconds MVonRX->MVonTX: 102.36 seconds DASonTX->MVonRX: 32.48 seconds
Plot single image
img{5}.plot(figure(123), label{5}, dynamic_range, compression);

Plot Figure - All 5 images
figure(3);
img{1}.plot(subplot(2,3,1), label{1}, dynamic_range, compression);
ax(1) = gca;
img{2}.plot(subplot(2,3,2), label{2}, dynamic_range, compression);
ax(2) = gca;
img{3}.plot(subplot(2,3,3), label{3}, dynamic_range, compression);
ax(3) = gca;
img{4}.plot(subplot(2,3,4), label{4}, dynamic_range, compression);
ax(4) = gca;
img{5}.plot(subplot(2,3,5), label{5}, dynamic_range, compression);
ax(5) = gca;
linkaxes(ax);
set(gcf,'Position',[29 33 1109 715]);

Plot timing bar chart
figure(4);
bar([time{:}]/60);
xticklabels(label);
xtickangle(45);
ylabel('Computation time (m)');
set(gca,'FontSize',14);
set(gcf,'Position',[29 33 800 400]);


Save figures
folder_path = 'figures'; mkdir(folder_path); b_data_compare = uff.beamformed_data(img{1}); b_data_compare.data(:,1,1,1) = img{1}.data; close all; for i = 1:length(img) b_data_compare.data(:,1,1,i) = img{i}.data./max(img{i}.data); f = figure(i); clf; img{i}.plot(f, [], dynamic_range, compression); set(gca, 'FontSize', 18); xlabel('x [mm]', 'FontSize', 20); ylabel('z [mm]', 'FontSize', 20); colorbar off; cb = colorbar; ylabel(cb, 'dB', 'FontSize', 18); set(cb, 'FontSize', 16); % Reduce whitespace by setting paper size to match figure set(f, 'PaperPositionMode', 'auto'); f.PaperUnits = 'inches'; f.PaperPosition = [0 0 6 5]; print(f, [folder_path, filesep, strrep(label{i}, '->', '_'), '_redone'], '-depsc2'); axis([0 10 20 28]); print(f, [folder_path, filesep, strrep(label{i}, '->', '_'), '_zoomed_redone'], '-depsc2'); end
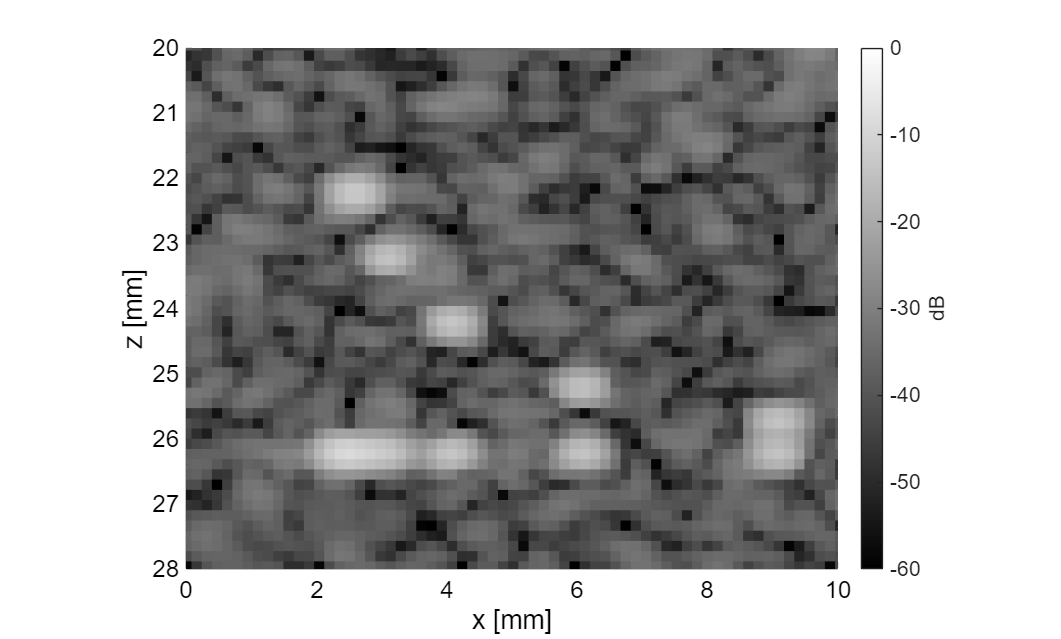





Compare beamformed data
b_data_compare.plot();

Plot timing bar chart
running_time_in_m = [time{1:5}]/60;
f99 = figure(100); clf;
bar([time{1:5}]/60);
labels_latex{1} = '$b^{\overline{T_{x DAS}}~\overline{R_{x DAS}}}$';
labels_latex{2} = '$b^{\overline{T_{x DAS}}~\overline{R_{x MV}}}$';
labels_latex{3} = '$b^{\overline{T_{x MV}}~\overline{R_{x DAS}}}$';
labels_latex{4} = '$b^{\overline{T_{x MV}}~\overline{R_{x MV}}}$';
labels_latex{5} = '$b^{\overline{R_{x MV}}~\overline{T_{x DAS}}}$';
%labels_latex{6} = '$b^{\overline{R_{x MV}}~\overline{T_{x DAS}}} \times {CF}^{0.2}$';
ylim([0 max(running_time_in_m)+1]);
ylabel('Computation time (m)');
set(gca,'FontSize',18);
x_pos = [1 2 3 4 5];
text(x_pos, running_time_in_m, num2str(running_time_in_m(:),'%.2f'), ...
'HorizontalAlignment','center', 'VerticalAlignment','bottom', 'FontSize',18);
set(gca, 'XTickLabel', labels_latex, 'TickLabelInterpreter', 'latex', 'FontSize',24);
set(gcf,'Position',[250 344 781 470]); grid on;
saveas(f99, [folder_path, filesep, 'adaptive_timing_redone'], 'eps2c');

=========================================================================
gCNR Analysis
=========================================================================
Define ROI positions (adjust for your data!)
xc_cyst = -8e-3; % cyst center x [m] zc_cyst = 24.25e-3; % cyst center z [m] xc_speckle = -1e-3; % speckle center x [m] zc_speckle = 30e-3; % speckle center z [m] radius = 3.5e-3; % radius [m] % Measure gCNR for all beamformers GCNR = zeros(1, 5); for i = 1:5 [~, ~, GCNR(i)] = tools.measure_contrast_circles(img{i}, xc_cyst, zc_cyst, xc_speckle, zc_speckle, radius, 0); end % Display results fprintf('\n=== gCNR Comparison ===\n'); for i = 1:5 fprintf('%-25s gCNR = %.3f\n', label{i}, GCNR(i)); end % Plot image with ROIs indicated f200 = figure(200); clf; img_data = img{1}.get_image('log'); imagesc(scan.x_axis*1e3, scan.z_axis*1e3, img_data); colormap gray; caxis([-dynamic_range 0]); axis image; hold on; % Draw cyst ROI (red) theta = linspace(0, 2*pi, 100); plot(xc_cyst*1e3 + radius*1e3*cos(theta), zc_cyst*1e3 + radius*1e3*sin(theta), 'r-', 'LineWidth', 2); % Draw speckle ROI (green) plot(xc_speckle*1e3 + radius*1e3*cos(theta), zc_speckle*1e3 + radius*1e3*sin(theta), 'g-', 'LineWidth', 2); hold off; set(gca, 'FontSize', 18); xlabel('x [mm]', 'FontSize', 20); ylabel('z [mm]', 'FontSize', 20); colorbar off; cb = colorbar; ylabel(cb, 'dB', 'FontSize', 18); set(cb, 'FontSize', 16); % Save with same settings as other images set(f200, 'PaperPositionMode', 'auto'); f200.PaperUnits = 'inches'; f200.PaperPosition = [0 0 6 5]; print(f200, [folder_path, filesep, 'DASonRX_DASonTX_ROI_locations_redone'], '-depsc2'); % gCNR bar plot f201 = figure(201); clf; bar(GCNR); xticklabels(label(1:5)); xtickangle(45); ylabel('gCNR'); title('Generalized Contrast-to-Noise Ratio Comparison'); ylim([0 1.1]); set(gca, 'FontSize', 14); grid on; % Add value labels on top of bars x_pos = 1:5; text(x_pos, GCNR, num2str(GCNR(:),'%.3f'), ... 'HorizontalAlignment','center', 'VerticalAlignment','bottom', 'FontSize', 12); saveas(f201, [folder_path, filesep, 'gCNR_res'], 'eps2c');
mean_background =
single
0.0615
mean_ROI =
single
8.4563e-04
CR =
single
-18.6162
CNR =
single
1.0911
mean_background =
0.0476
mean_ROI =
4.1221e-04
CR =
-20.6271
CNR =
1.0767
mean_background =
0.0615
mean_ROI =
8.4563e-04
CR =
-18.6162
CNR =
1.0911
mean_background =
0.0476
mean_ROI =
4.1221e-04
CR =
-20.6271
CNR =
1.0767
mean_background =
0.0666
mean_ROI =
7.4225e-04
CR =
-19.5313
CNR =
1.0922
=== gCNR Comparison ===
DASonRX->DASonTX gCNR = 0.928
MVonRX->DASonTX gCNR = 0.948
DASonRX->MVonTX gCNR = 0.928
MVonRX->MVonTX gCNR = 0.948
DASonTX->MVonRX gCNR = 0.940
Warning: Exported image displays axes toolbar. To remove axes toolbar from
image, export again.


=========================================================================
FWHM / 6dB Resolution Analysis
=========================================================================
Define point target location (adjust for your data!) PICMUS numerical phantom has point targets - find the one you want to analyze
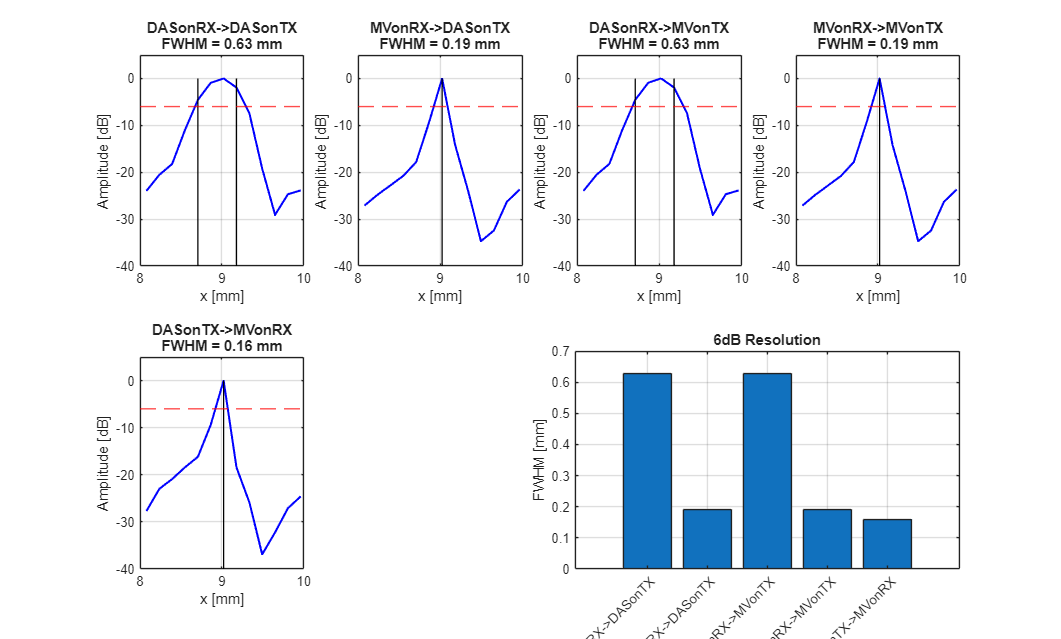
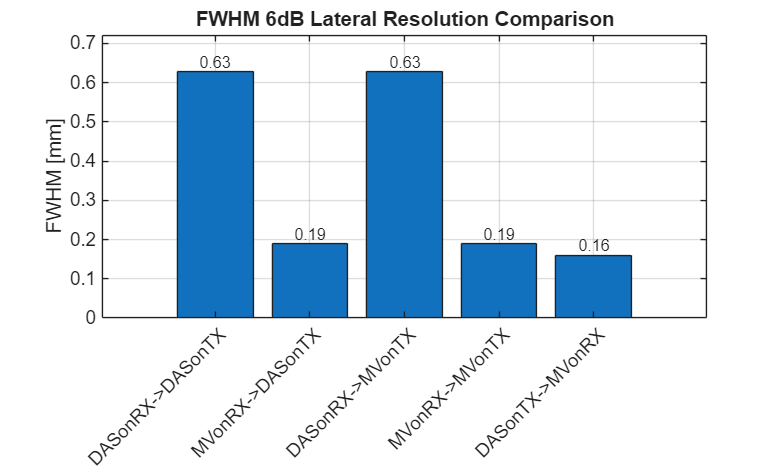
z_target = 26.2e-3; % target depth in meters x_target = 9e-3; % target lateral position in meters (0 = center) x_range = 1e-3; % range around target to analyze [m] % Find the z-index of the point target [~, z_idx] = min(abs(scan.z_axis - z_target)); % Find x indices for the analysis range x_idx_min = find(scan.x_axis >= (x_target - x_range), 1, 'first'); x_idx_max = find(scan.x_axis <= (x_target + x_range), 1, 'last'); x_range_idx = x_idx_min:x_idx_max; % Measure FWHM for all 6 beamformers with plots figure(300); clf; res = zeros(1, 5); for m = 1:5%6 % Get lateral profile at target depth (in dB) img_dB = img{m}.get_image('log'); lateral_full = img_dB(z_idx, :); % Extract region around target x_mm = scan.x_axis(x_range_idx) * 1e3; lateral = lateral_full(x_range_idx); lateral = lateral - max(lateral); % Normalize to 0 dB at peak % Calculate 6dB resolution (no internal plotting) res(m) = compute_fwhm_6dB(x_mm, lateral); % Plot lateral profile subplot(2, 4, m); plot(x_mm, lateral, 'b-', 'LineWidth', 1.5); hold on; % Draw -6dB line yline(-6, 'r--', 'LineWidth', 1); % Find and mark -6dB crossing points idx_above = find(lateral >= -6); if ~isempty(idx_above) x_left = x_mm(idx_above(1)); x_right = x_mm(idx_above(end)); plot([x_left x_left], [-60 0], 'k-', 'LineWidth', 1); plot([x_right x_right], [-60 0], 'k-', 'LineWidth', 1); end hold off; xlabel('x [mm]'); ylabel('Amplitude [dB]'); title(sprintf('%s\nFWHM = %.2f mm', label{m}, res(m))); ylim([-40 5]); grid on; set(gca, 'FontSize', 10); end % Add summary subplot subplot(2, 4, [7 8]); bar(res); xticklabels(label); xtickangle(45); ylabel('FWHM [mm]'); title('6dB Resolution'); set(gca, 'FontSize', 10); grid on; % Display results fprintf('\n=== FWHM (6dB Resolution) Comparison ===\n'); for i = 1:5 fprintf('%-25s FWHM = %.3f mm\n', label{i}, res(i)); end % Resolution bar plot f301 = figure(301); clf; bar(res); xticklabels(label(1:5)); xtickangle(45); ylabel('FWHM [mm]'); title('FWHM 6dB Lateral Resolution Comparison'); set(gca, 'FontSize', 14); grid on; set(gcf,'Position',[250 344 781 470]); % Add value labels on top of bars x_pos = 1:5; text(x_pos, res, num2str(res(:),'%.2f'), ... 'HorizontalAlignment','center', 'VerticalAlignment','bottom', 'FontSize', 12); ylim([0 max(res)*1.15]); saveas(f301, [folder_path, filesep, 'FWHM_res'], 'eps2c');
=========================================================================
Lateral Profile Plot
=========================================================================
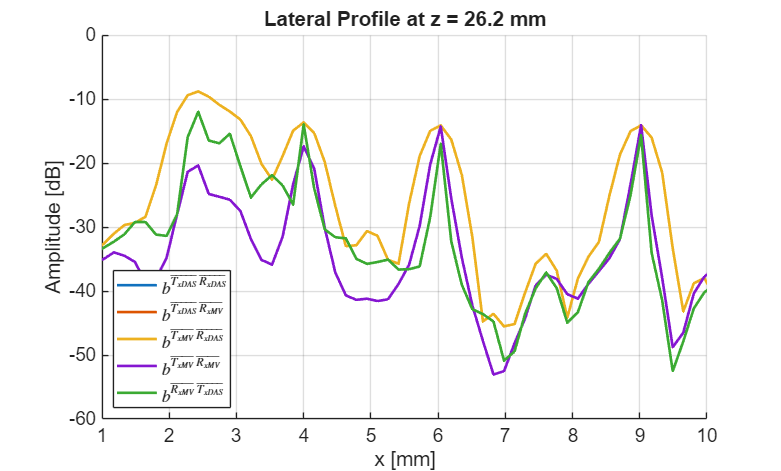
f400 = figure(400); clf; % Get lateral profiles at target depth colors = lines(5); hold on; for i = 1:5 img_dB = img{i}.get_image('log'); lateral = img_dB(z_idx, :); plot(scan.x_axis*1e3, lateral, 'LineWidth', 2, 'Color', colors(i,:)); end hold off; xlabel('x [mm]'); ylabel('Amplitude [dB]'); title(sprintf('Lateral Profile at z = %.1f mm', z_target*1e3)); legend(labels_latex(1:5), 'Location', 'sw', 'Interpreter', 'latex'); ylim([-dynamic_range 0]); xlim([1 10]); grid on; set(gca, 'FontSize', 14); set(gcf,'Position',[250 344 781 470]); grid on; saveas(f400, [folder_path, filesep, 'lateral_profile_plot'], 'eps2c');
=========================================================================
Combined Bar Charts Figure
=========================================================================
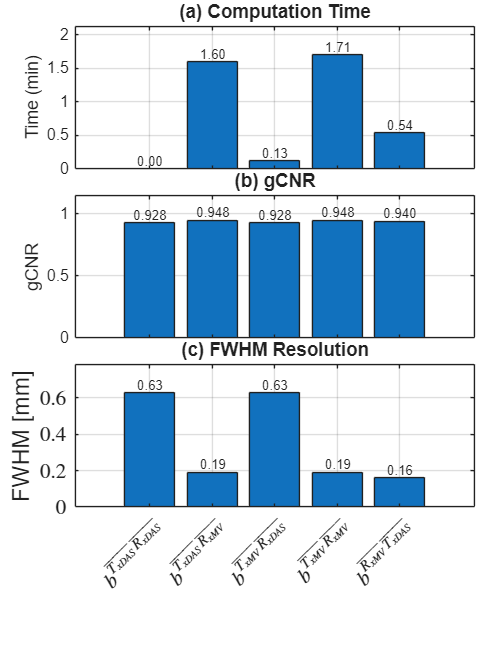
f500 = figure(500); clf; set(gcf, 'Position', [100 100 500 650]); % Manually position subplots for tighter layout [left bottom width height] % Subplot 1: Computation Time (top) ax1 = axes('Position', [0.15 0.74 0.80 0.22]); bar(running_time_in_m); ylabel('Time (min)'); ylim([0 max(running_time_in_m)*1.25]); set(gca, 'FontSize', 12); set(gca, 'XTickLabel', []); grid on; x_pos = 1:length(running_time_in_m); text(x_pos, running_time_in_m, num2str(running_time_in_m(:),'%.2f'), ... 'HorizontalAlignment','center', 'VerticalAlignment','bottom', 'FontSize', 10); title('(a) Computation Time', 'FontSize', 14); % Subplot 2: gCNR (middle) ax2 = axes('Position', [0.15 0.48 0.80 0.22]); bar(GCNR); ylabel('gCNR'); ylim([0 1.15]); set(gca, 'FontSize', 12); set(gca, 'XTickLabel', []); grid on; x_pos = 1:5; text(x_pos, GCNR, num2str(GCNR(:),'%.3f'), ... 'HorizontalAlignment','center', 'VerticalAlignment','bottom', 'FontSize', 10); title('(b) gCNR', 'FontSize', 14); % Subplot 3: FWHM Resolution (bottom) ax3 = axes('Position', [0.15 0.22 0.80 0.22]); bar(res); ylabel('FWHM [mm]'); ylim([0 max(res)*1.25]); set(gca, 'FontSize', 12); set(gca, 'XTickLabel', labels_latex(1:5), 'TickLabelInterpreter', 'latex', 'XTickLabelRotation', 45, 'FontSize', 16); grid on; x_pos = 1:5; text(x_pos, res, num2str(res(:),'%.2f'), ... 'HorizontalAlignment','center', 'VerticalAlignment','bottom', 'FontSize', 10); title('(c) FWHM Resolution', 'FontSize', 14); % Save combined figure saveas(f500, [folder_path, filesep, 'combined_metrics_redone'], 'eps2c');
=========================================================================
Summary Table
=========================================================================
fprintf('\n=== Summary ===\n'); fprintf('%-25s %10s %10s %10s\n', 'Method', 'Time (s)', 'gCNR', 'FWHM (mm)'); fprintf('%s\n', repmat('-', 1, 60)); for i = 1:5 fprintf('%-25s %10.2f %10.3f %10.3f\n', label{i}, time{i}, GCNR(i), res(i)); end
=== Summary === Method Time (s) gCNR FWHM (mm) ------------------------------------------------------------ DASonRX->DASonTX 0.09 0.928 0.627 MVonRX->DASonTX 95.94 0.948 0.190 DASonRX->MVonTX 7.55 0.928 0.627 MVonRX->MVonTX 102.36 0.948 0.190 DASonTX->MVonRX 32.48 0.940 0.161
=========================================================================
Local Functions
=========================================================================
function res = compute_fwhm_6dB(x_axis, y_signal) % COMPUTE_FWHM_6DB Compute the -6dB width (FWHM) of a signal % x_axis: lateral position in mm % y_signal: signal in dB (should be normalized so max = 0) % res: -6dB width in mm % Interpolate for better resolution coeff = 10; x_interp = linspace(x_axis(1), x_axis(end), length(x_axis) * coeff); y_interp = interp1(x_axis, y_signal, x_interp, 'spline'); % Find points above -6dB idx_above = find(y_interp >= -6); if isempty(idx_above) res = NaN; warning('Could not find -6dB crossing points'); return; end % Calculate width res = x_interp(idx_above(end)) - x_interp(idx_above(1)); end
=== FWHM (6dB Resolution) Comparison === DASonRX->DASonTX FWHM = 0.627 mm MVonRX->DASonTX FWHM = 0.190 mm DASonRX->MVonTX FWHM = 0.627 mm MVonRX->MVonTX FWHM = 0.190 mm DASonTX->MVonRX FWHM = 0.161 mm