Resolution of Delay Multiply And Sum on FI data from an UFF file
by Ole Marius Hoel Rindal olemarius@olemarius.net 28.05.2017
Contents
Setting up file path
To read data from a UFF file the first we need is, you guessed it, a UFF file. We check if it is on the current path and download it from the USTB websever.
clear all; close all; % data location url='http://ustb.no/datasets/'; % if not found downloaded from here local_path = [ustb_path(),'/data/']; % location of example data % Choose dataset filename='Alpinion_L3-8_FI_hyperechoic_scatterers.uff'; % check if the file is available in the local path or downloads otherwise tools.download(filename, url, local_path);
Reading channel data from UFF file
channel_data=uff.read_object([local_path filename],'/channel_data'); % Check that the user have the correct version of the dataset if(strcmp(channel_data.version{1},'1.0.2')~=1) error(['Wrong version of the dataset. Please delete ',local_path,... filename,' and rerun script.']); end
UFF: reading channel_data [uff.channel_data] UFF: reading sequence [uff.wave] [====================] 100%
%Print info about the dataset
channel_data.print_authorship
Name: FI dataset of hyperechoic cyst and points scatterers recorded on an Alpinion scanner with a L3-8 Probe from a CIRS General Purpose Ultrasound Phantom Reference: www.ultrasoundtoolbox.com Author(s): Ole Marius Hoel Rindal <olemarius@olemarius.net> Muyinatu Lediju Bell <mledijubell@jhu.edu> Version: 1.0.2
Define Scan
Define the image coordinates we want to beamform in the scan object. Notice that we need to use quite a lot of samples in the z-direction. This is because the DMAS creates an "artificial" second harmonic signal, so we need high enough sampling frequency in the image to get a second harmonic signal.
z_axis=linspace(25e-3,45e-3,1024).'; x_axis=zeros(channel_data.N_waves,1); for n=1:channel_data.N_waves x_axis(n) = channel_data.sequence(n).source.x; end scan=uff.linear_scan('x_axis',x_axis,'z_axis',z_axis);
Set up the processing pipeline
pipe=pipeline(); pipe.channel_data=channel_data; pipe.scan=scan; pipe.transmit_apodization.window=uff.window.scanline; pipe.receive_apodization.window=uff.window.none; pipe.receive_apodization.f_number=1.7;
Define the DAS beamformer
das = midprocess.das();
%Sum only on transmit, so that we can do DMAS on receice
das.dimension = dimension.transmit();
Create the DMAS image using the delay_multiply_and_sum postprocess
dmas = postprocess.delay_multiply_and_sum();
dmas.dimension = dimension.receive;
dmas.channel_data = channel_data;
dmas.receive_apodization = pipe.receive_apodization;
b_data_dmas=pipe.go({das dmas});
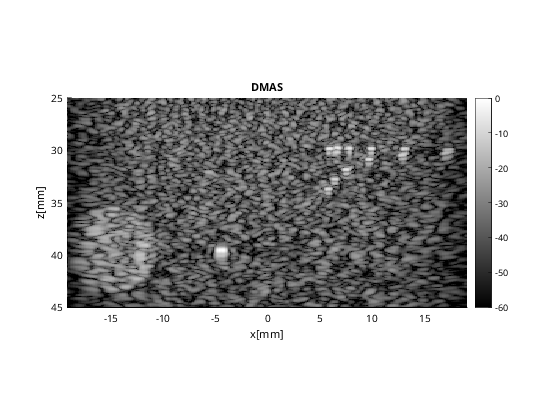
% beamforming
b_data_dmas.plot(100,'DMAS');
USTB MEX C beamformer...Completed in 0.42 seconds.
f_stop =
single
10913708
Warning: If the result looks funky, you might need to tune the filter paramters
of DMAS using the filter_freqs property. Use the plot to check that everything
is OK.

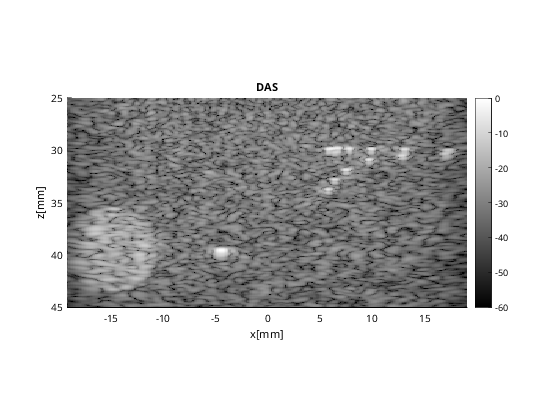
Beamform DAS image
Notice that I redefine the beamformer to summing on both transmit and receive.
das.dimension = dimension.both();
b_data_das=pipe.go({das});
b_data_das.plot([],'DAS');
USTB MEX C beamformer...Completed in 0.40 seconds.

Plot both images in same plot
Plot both in same plot with connected axes, try to zoom!
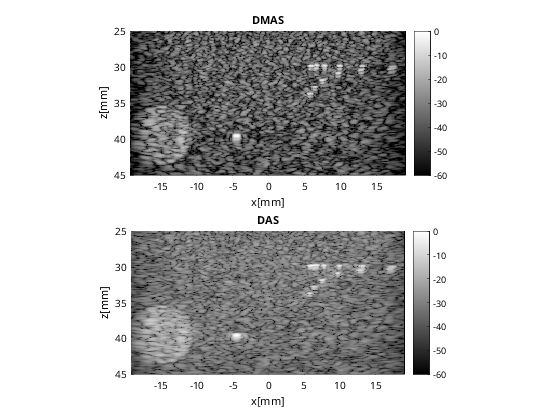
f3 = figure(3);clf b_data_dmas.plot(subplot(2,1,1),'DMAS'); % Display image ax(1) = gca; b_data_das.plot(subplot(2,1,2),'DAS'); % Display image ax(2) = gca; linkaxes(ax);
Compare resolution
Plot the lateral line through some of the scatterers
% Let's get the images as a N_z_axis x N_x_axis image
dmas_img = b_data_dmas.get_image();
das_img = b_data_das.get_image();
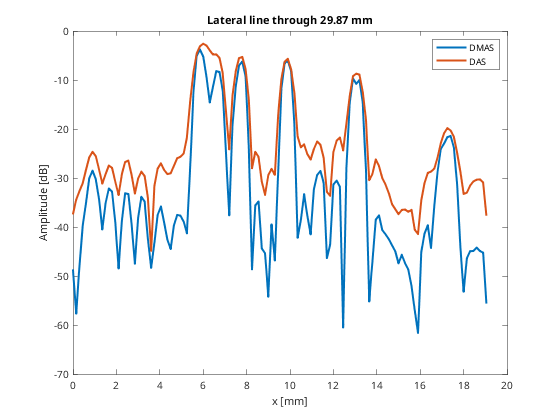
So that we can plot the line through the group of scatterers
line_idx = 250; figure(4);clf; plot(b_data_dmas.scan.x_axis*10^3,dmas_img(line_idx,:),... 'DisplayName','DMAS','LineWidth',2);hold on; plot(b_data_das.scan.x_axis*10^3,das_img(line_idx,:),... 'DisplayName','DAS','LineWidth',2); xlabel('x [mm]');xlim([0 20]);ylabel('Amplitude [dB]');legend show title(sprintf('Lateral line through %.2f mm',... b_data_dmas.scan.z_axis(line_idx)*10^3));
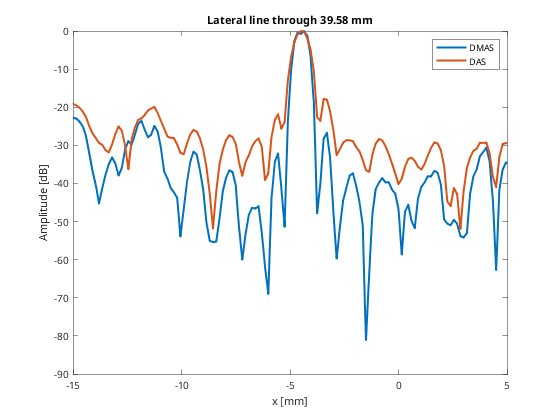
%So that we can plot the line through the aingle scatterer line_idx = 747; figure(5);clf; plot(b_data_dmas.scan.x_axis*10^3,dmas_img(line_idx,:),... 'DisplayName','DMAS','LineWidth',2);hold on; plot(b_data_das.scan.x_axis*10^3,das_img(line_idx,:),... 'DisplayName','DAS','LineWidth',2); xlabel('x [mm]');xlim([-15 5]);ylabel('Amplitude [dB]');legend show title(sprintf('Lateral line through %.2f mm',... b_data_dmas.scan.z_axis(line_idx)*10^3));